Exploring Relationships and Bivariate Analysis
Bard College | Introduction to Data Analytics
Bivariate analysis
Comparing distributions
Scatter plots
Calculate correlation coefficients
Correlation plots
Cross tabulations (
count(),group_byandsummarize)Bivariate mapping (later)
Covariance (later)
Simple linear regression (later)
Getting started
Before we get started, load the palmerpenguins package.
This package contains the dataset
penguinsUse
?penguinsto learn more about the data.
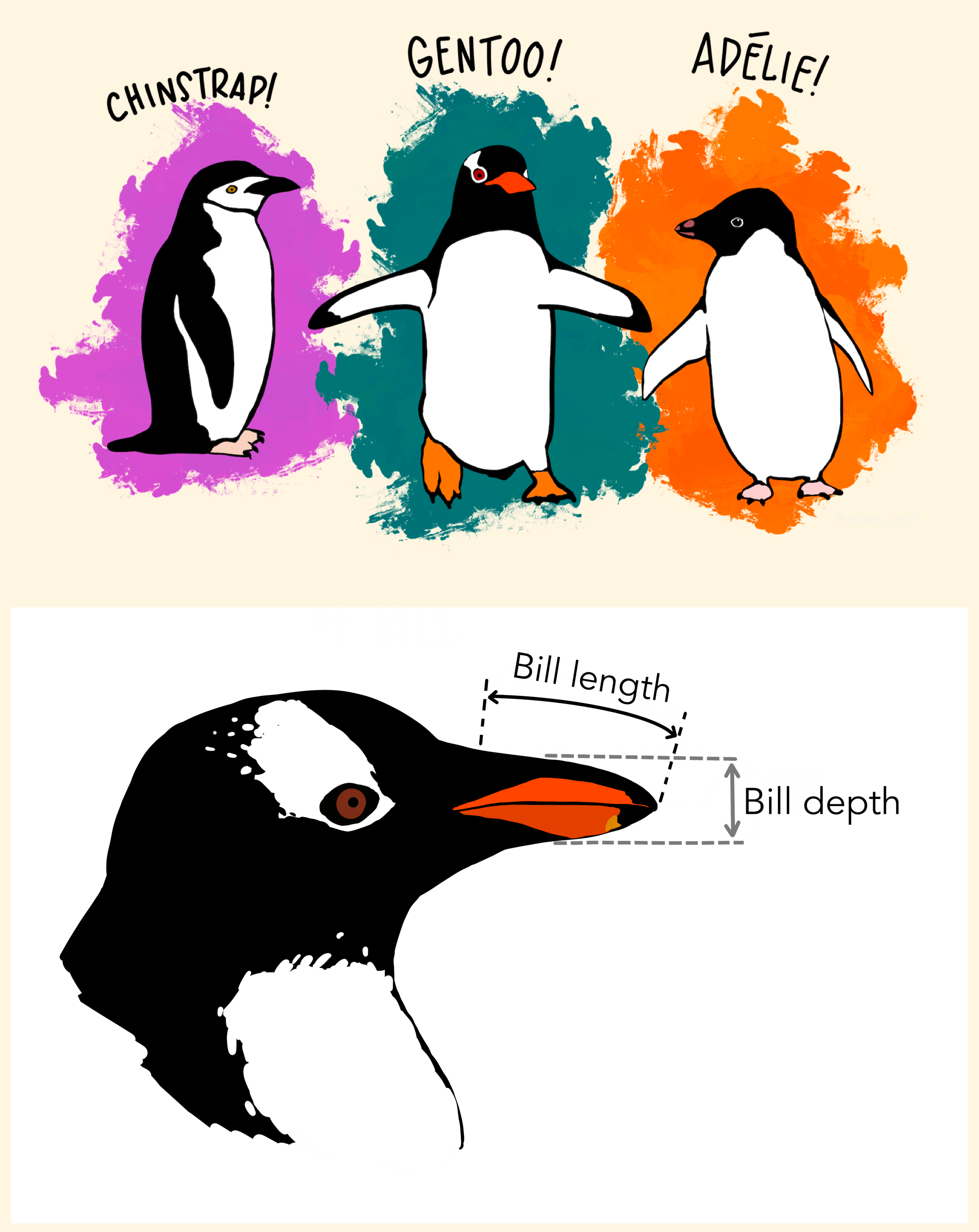
Data: Palmer Penguins
From an earlier session…
Measurements for penguin species, island in Palmer Archipelago, size (flipper length, body mass, bill dimensions), and sex.
Rows: 344
Columns: 8
$ species <fct> Adelie, Adelie, Adelie, Adelie, Adelie, Adelie, Adel…
$ island <fct> Torgersen, Torgersen, Torgersen, Torgersen, Torgerse…
$ bill_length_mm <dbl> 39.1, 39.5, 40.3, NA, 36.7, 39.3, 38.9, 39.2, 34.1, …
$ bill_depth_mm <dbl> 18.7, 17.4, 18.0, NA, 19.3, 20.6, 17.8, 19.6, 18.1, …
$ flipper_length_mm <int> 181, 186, 195, NA, 193, 190, 181, 195, 193, 190, 186…
$ body_mass_g <int> 3750, 3800, 3250, NA, 3450, 3650, 3625, 4675, 3475, …
$ sex <fct> male, female, female, NA, female, male, female, male…
$ year <int> 2007, 2007, 2007, 2007, 2007, 2007, 2007, 2007, 2007…Compare distributions
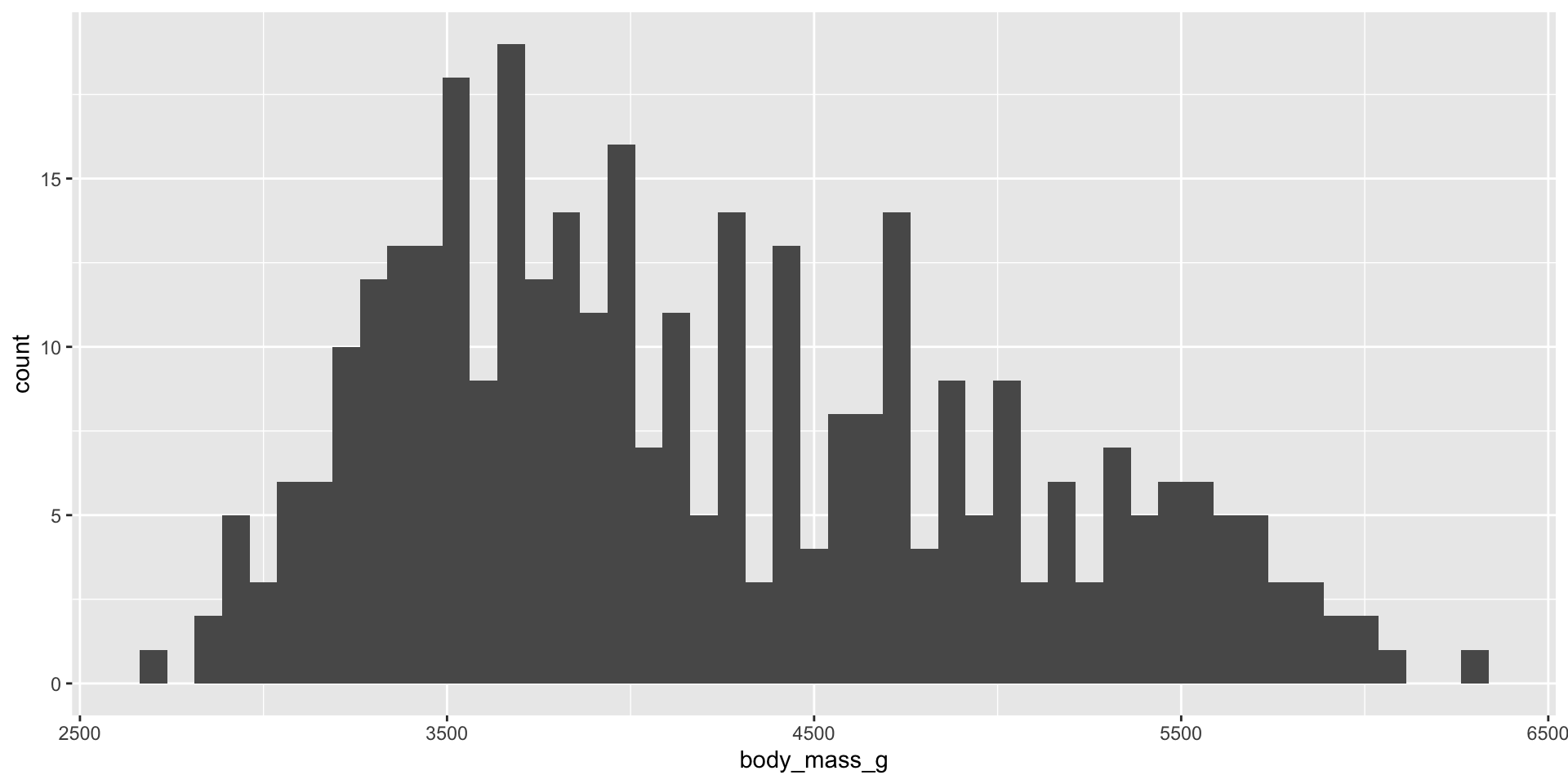
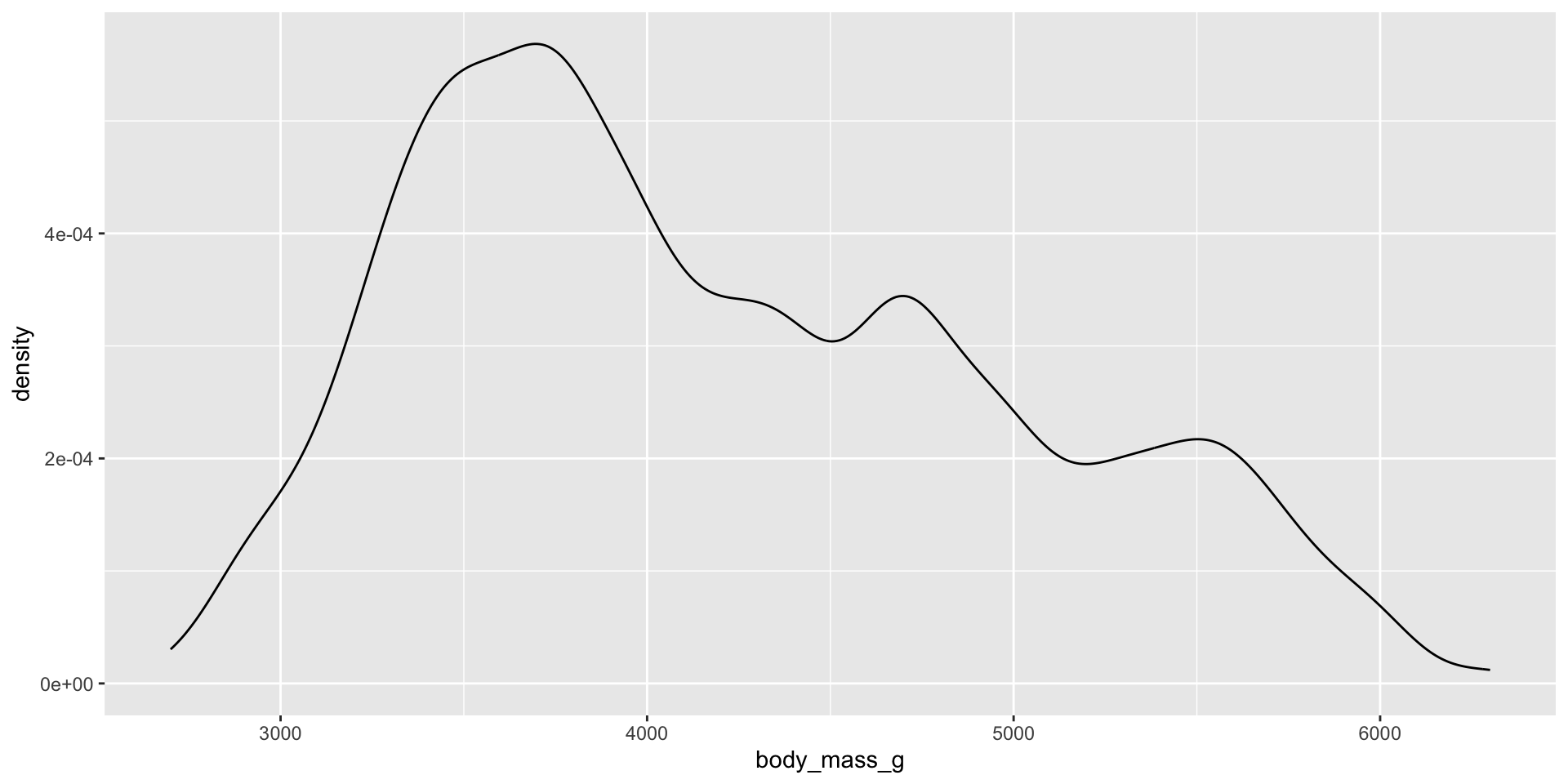
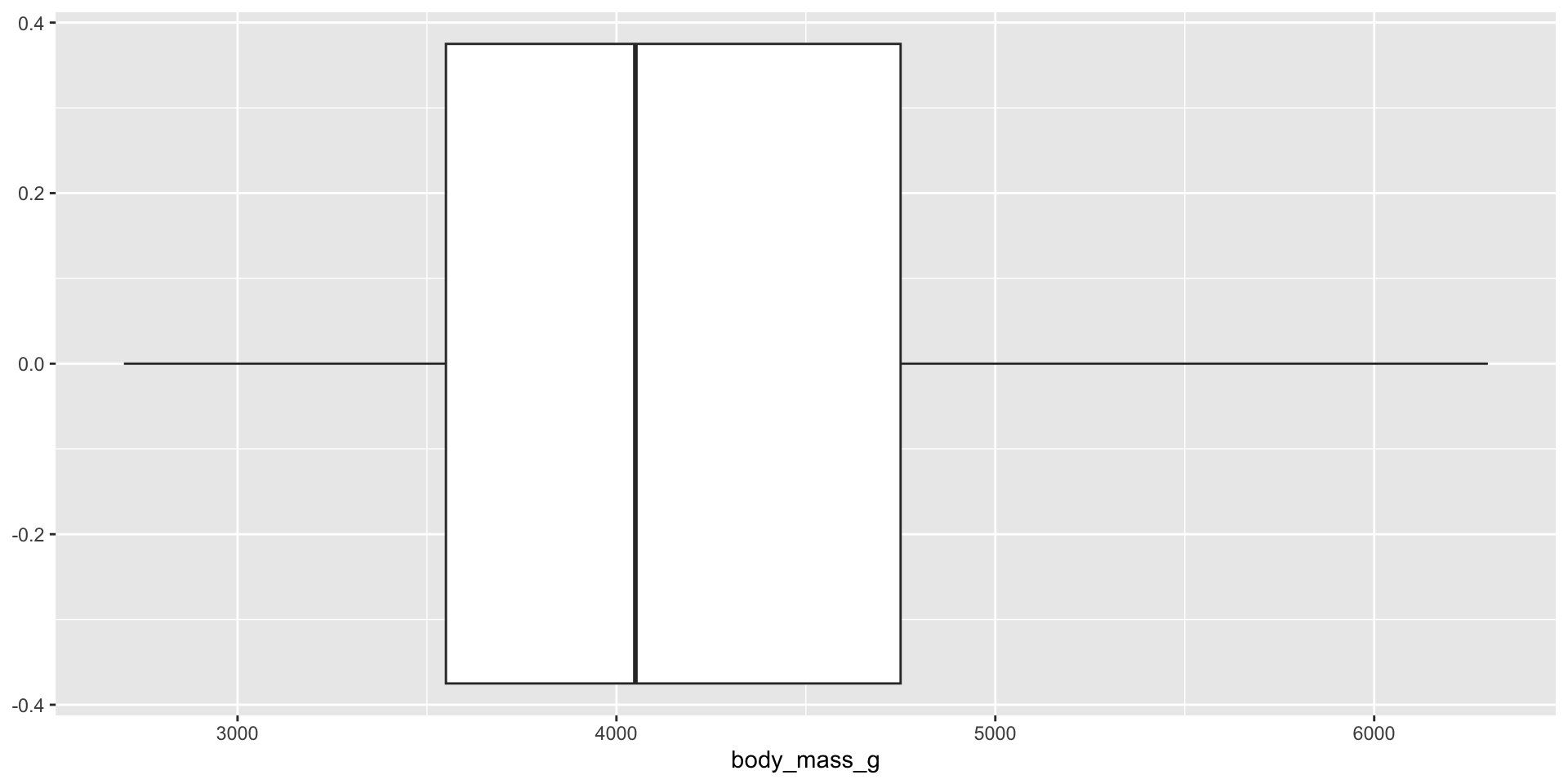
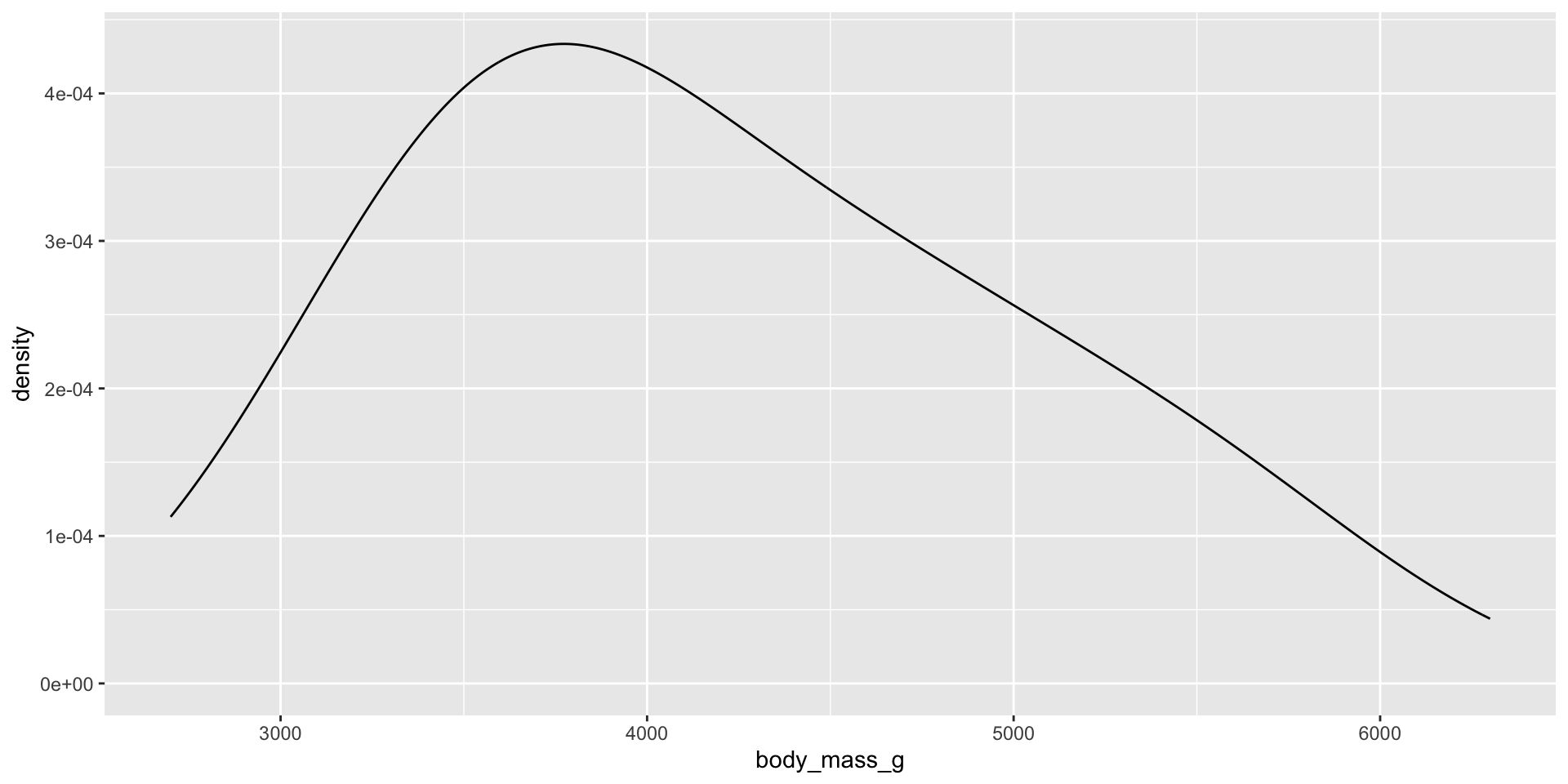
Measure and compare central tendency and spread
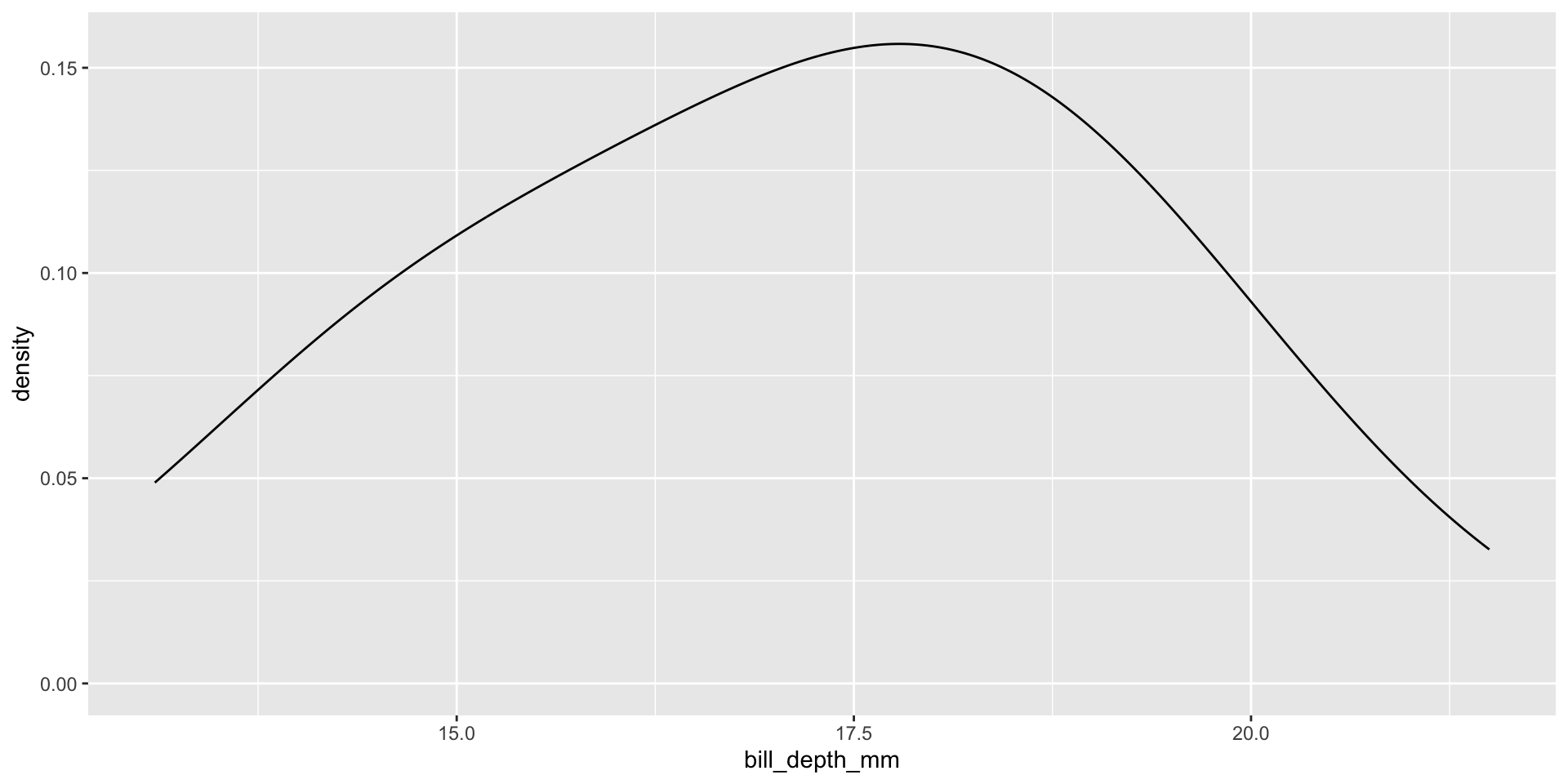
Histograms, density plots, and box plots
Reminders: Skew
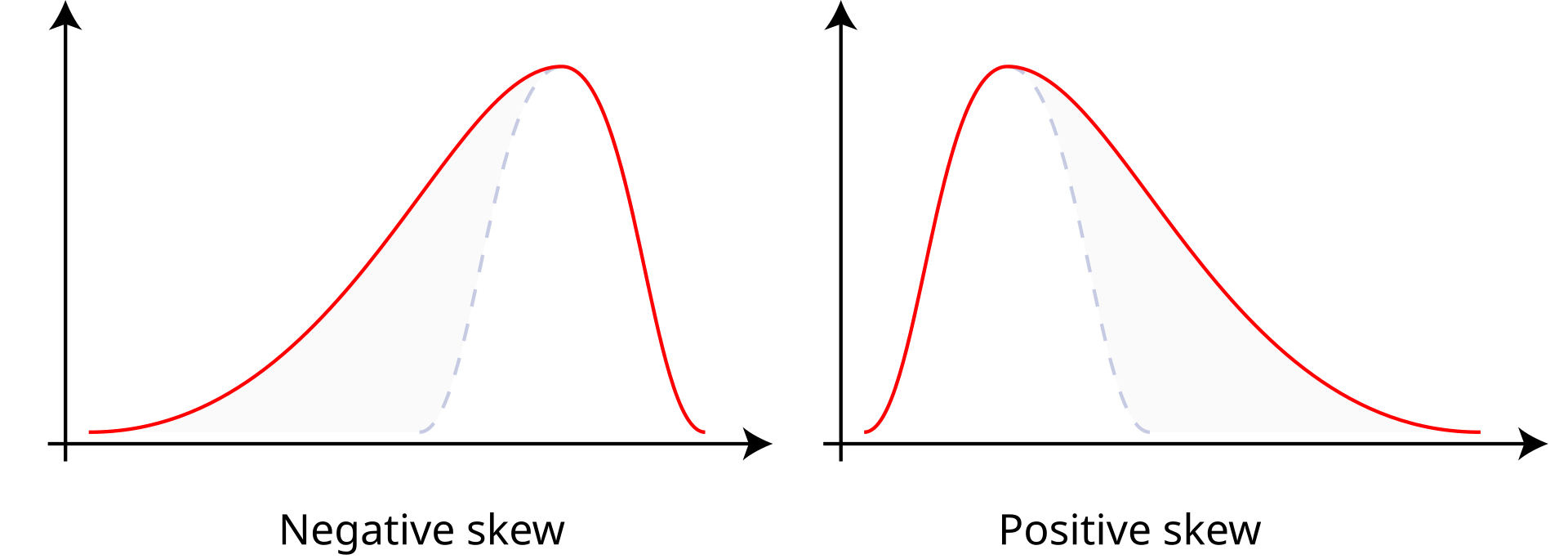
- Positive skew: The right tail is longer; the mass of the distribution is concentrated on the left of the figure. Right or positive skew refers to the right tail being drawn out and, often, the mean being skewed to the right of a typical center (median) of the data.
- Negative skew: The left tail is longer; the mass of the distribution is concentrated on the right of the figure. Left or negative skew refers to the left tail being drawn out and, often, the mean being skewed to the left of a typical center of the data.
Central tendency
For data from skewed distributions, the median is better than the mean because it isn’t influenced by extremely large values.
Which central tendency measure is most appropriate?
Measure spread or variability with standard deviation
Standard deviation
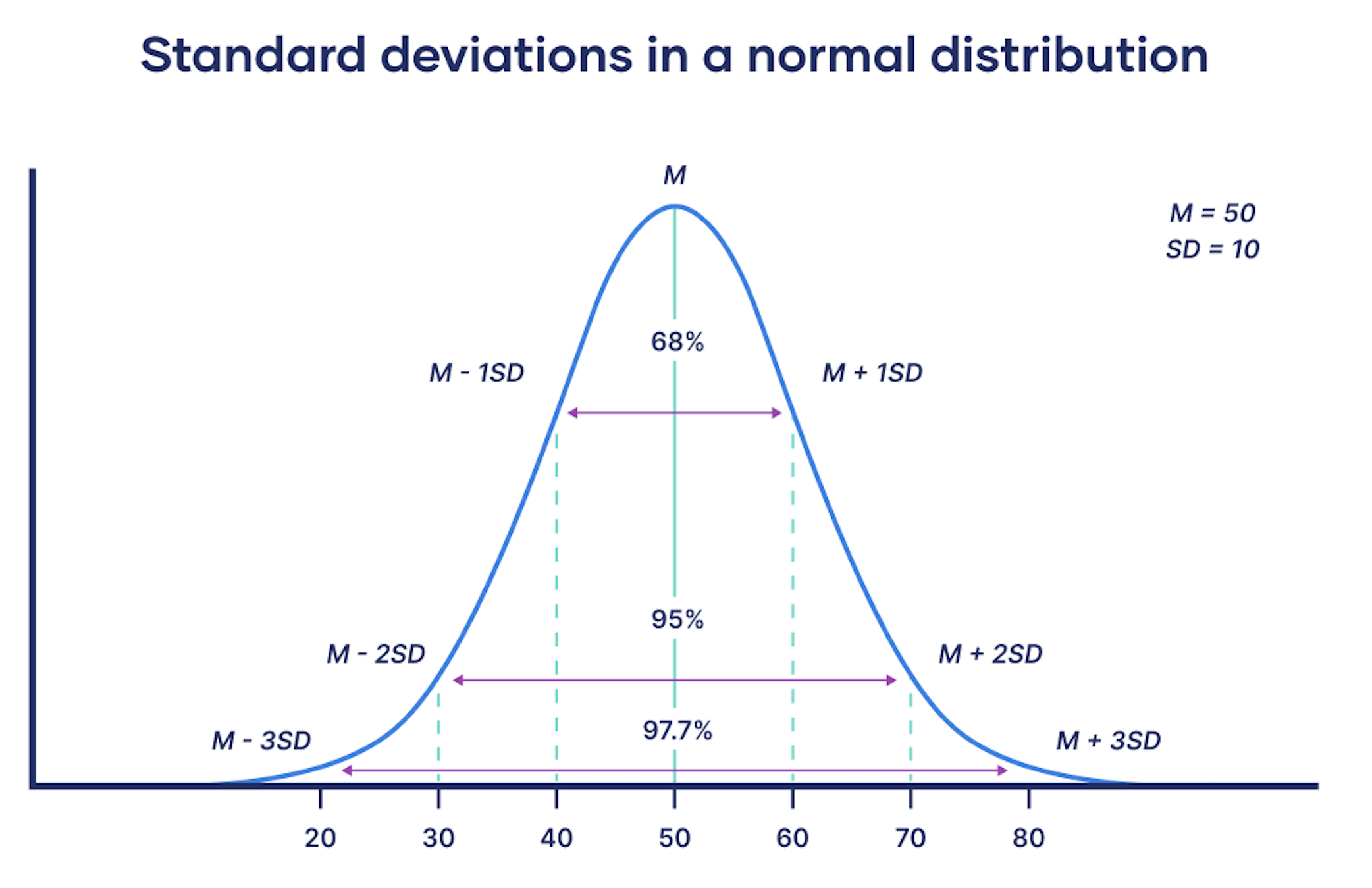
Standard deviation
For each value in the data set, we compute its deviation from the mean
\[ x_i - \bar{x} \]This will generate some negative and some positive values, so to get them to be positive we square them.
Then, add these up to compute the average deviation, or sample variance:
\[s^{2}= ∑(x_i−\bar{x})^2 / n-1\]
Take the square root to get the standard deviation: \[s\]
Calculating standard deviation
Let’s try this out in R
Calculating standard deviation
Standard deviation
Calculate standard deviation with dplyr functions
Calculate step-by-step
calc_standard_deviation <- df1 |>
mutate(deviation_from_mean = nums - mean(nums),
deviation_squared = deviation_from_mean^2) |>
summarize(sum_deviation_squared = sum(deviation_squared),
s2 = sum_deviation_squared/(17-1), # See formula for s-squared on previous slide
sd = sqrt(s2))
calc_standard_deviation sum_deviation_squared s2 sd
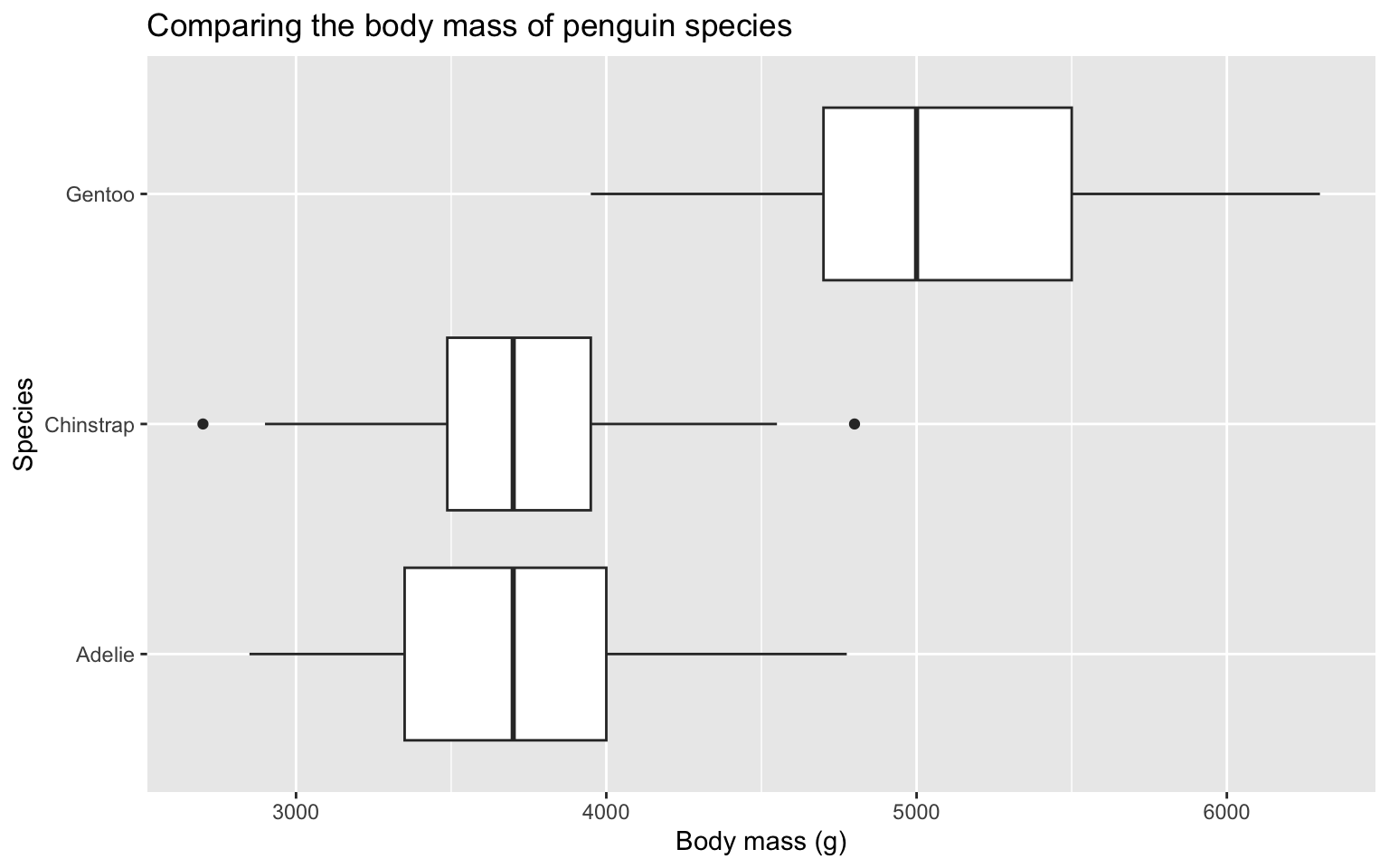
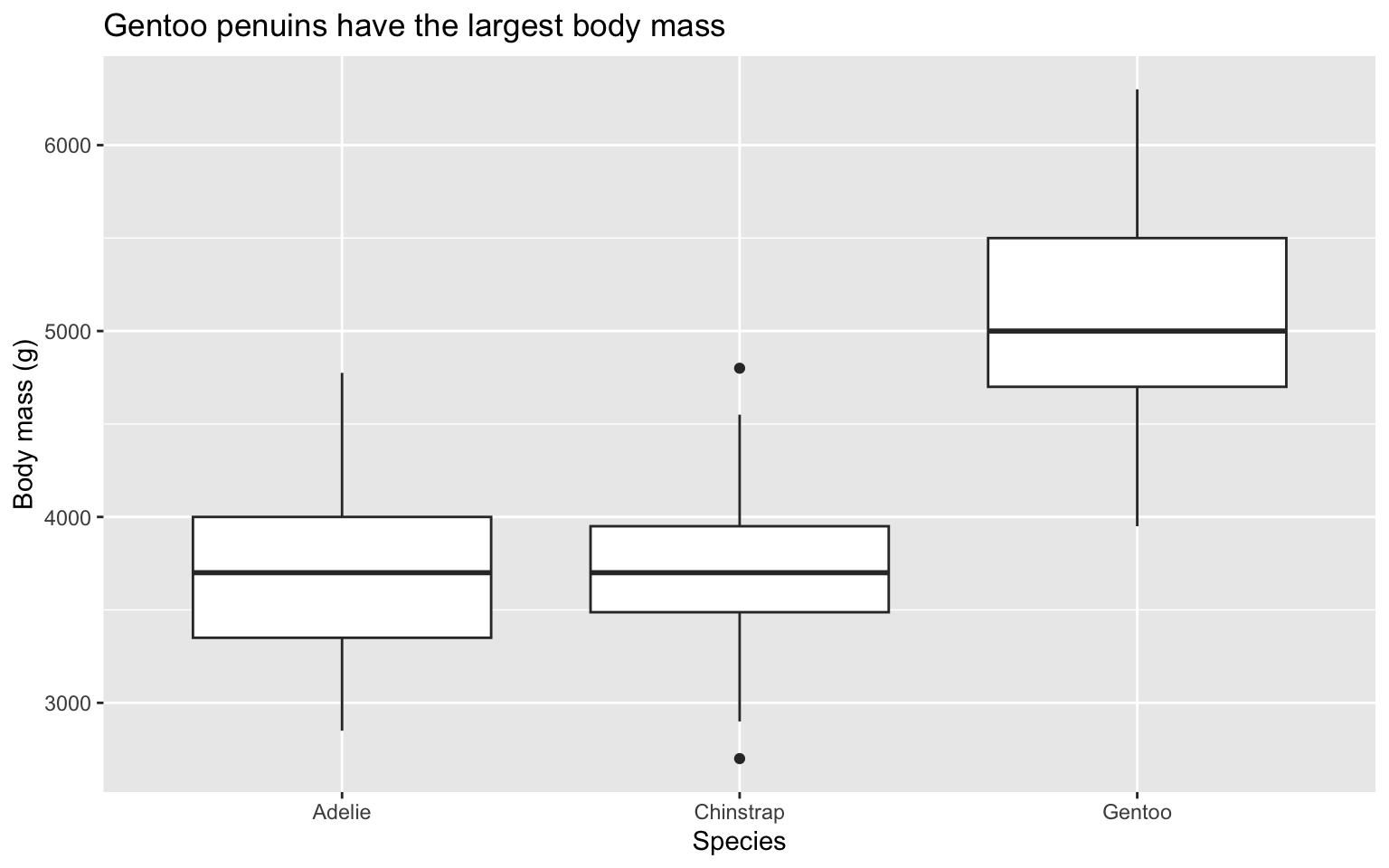
1 3672 229.5 15.14926Standard deviation of penguin body mass by species
Which species has the most variation in body mass (grams)?
Compare distributions with side-by-side box plots
Side-by-side box plots
Side-by-side box plots
Create a quick summary with summary
species island bill_length_mm bill_depth_mm
Adelie :152 Biscoe :168 Min. :32.10 Min. :13.10
Chinstrap: 68 Dream :124 1st Qu.:39.23 1st Qu.:15.60
Gentoo :124 Torgersen: 52 Median :44.45 Median :17.30
Mean :43.92 Mean :17.15
3rd Qu.:48.50 3rd Qu.:18.70
Max. :59.60 Max. :21.50
NA's :2 NA's :2
flipper_length_mm body_mass_g sex year
Min. :172.0 Min. :2700 female:165 Min. :2007
1st Qu.:190.0 1st Qu.:3550 male :168 1st Qu.:2007
Median :197.0 Median :4050 NA's : 11 Median :2008
Mean :200.9 Mean :4202 Mean :2008
3rd Qu.:213.0 3rd Qu.:4750 3rd Qu.:2009
Max. :231.0 Max. :6300 Max. :2009
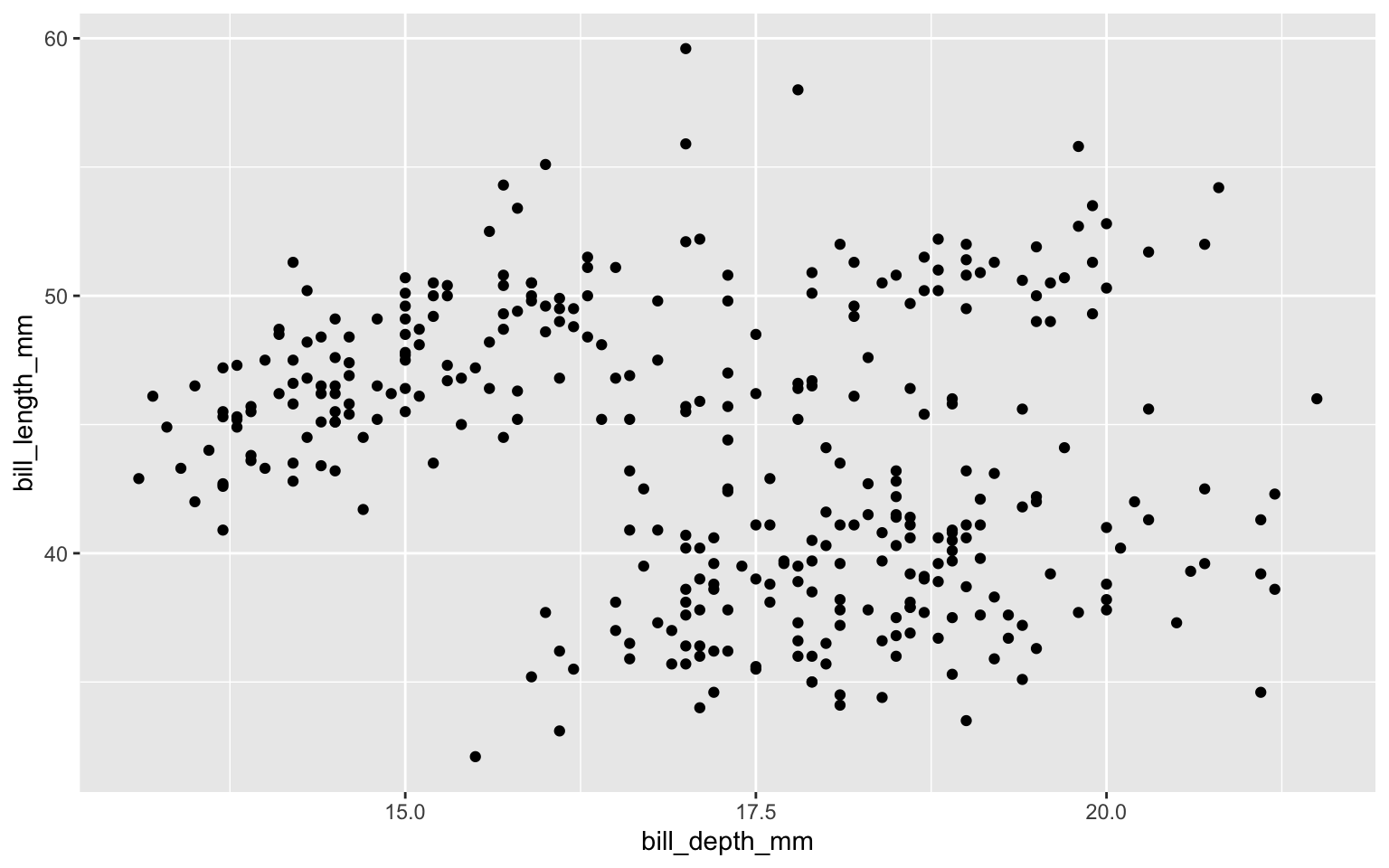
NA's :2 NA's :2 Show relationships between variables with scatter plots
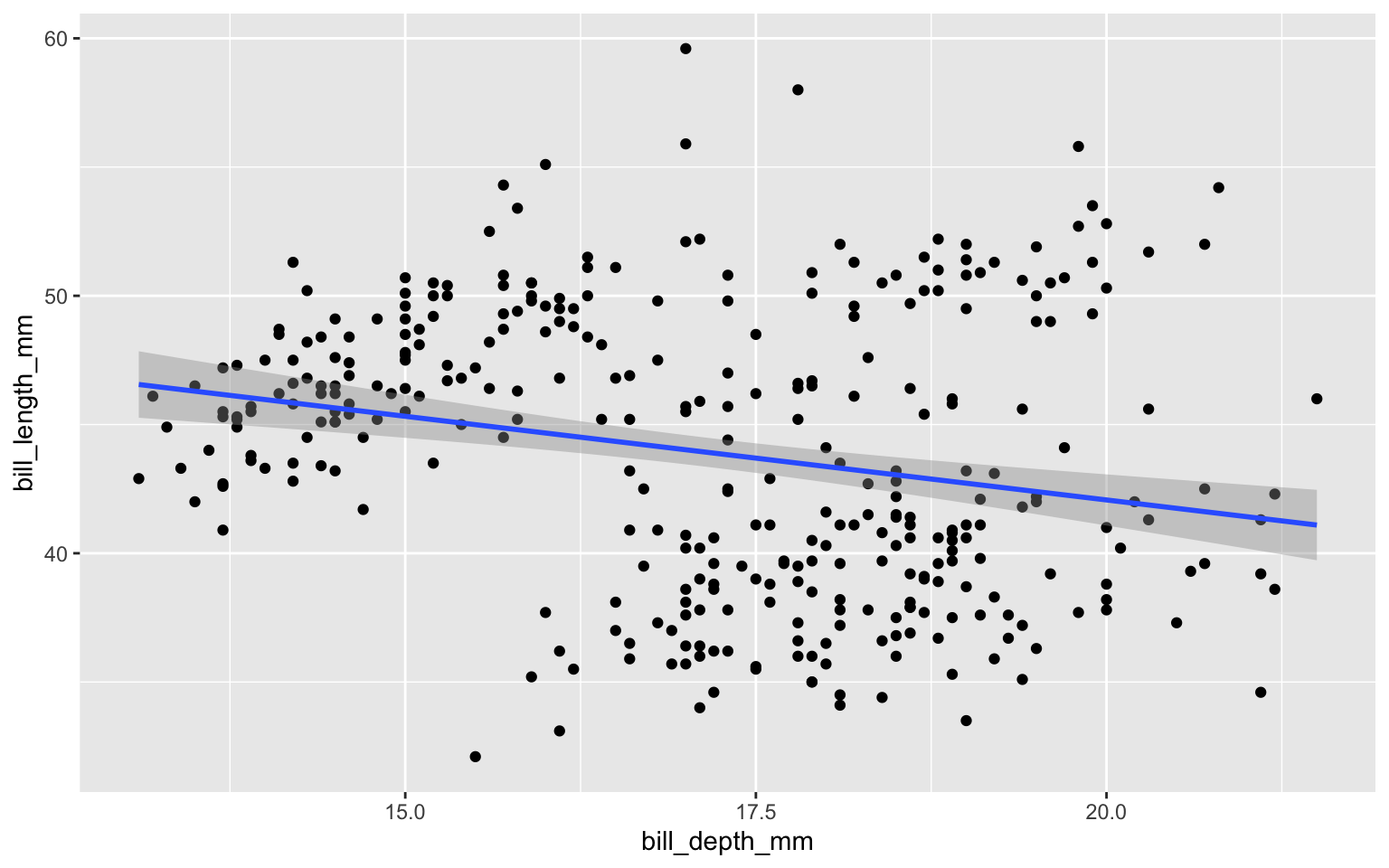
Scatter plots
Scatter plots: add linear regression line
Calculating correlation coefficients with cor()
Correlation coefficients
Use the
cor()function to calculate a Pearson’s R correlation coefficient (the default correlation coefficient type).Pearson R is used for linear relationships between two continuous, normally distributed variables.
-
The
cor()function comes from thestatspackage, which contains functions for statistical calculations and random number generation.- For a complete list of functions, use
library(help = "stats").
- For a complete list of functions, use
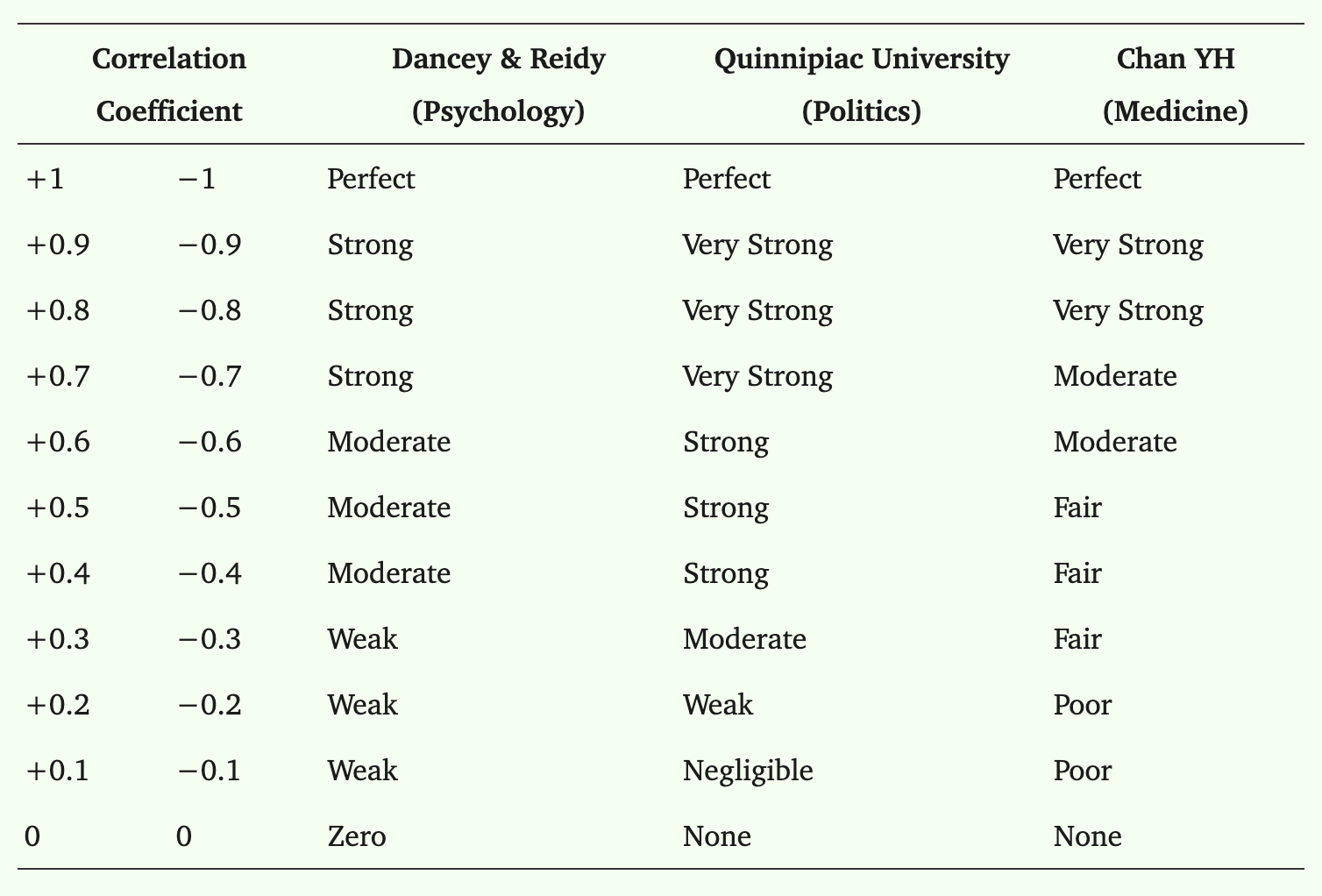
Correlation coefficients
Pearson’s R is one of the most popular correlation coefficients, and there are others that are useful based on the data types of the variables you are analyzing: numerical, ordinal, binary.
Guide to correlation coefficients: https://pmc.ncbi.nlm.nih.gov/articles/PMC6107969/
use = "complete.obs"tells cor() to only use rows where both variables are not missing (i.e., no NA values).
Scatter plots: add linear regression line and correlation coefficient
corr <- penguins |>
summarize(pearson_r = cor(bill_depth_mm, bill_length_mm, use = "complete.obs")) |>
pull(pearson_r) # pull() is similar to $ from base R. It's mostly useful because it works better in tidyverse pipes.
ggplot(penguins,
aes(x = bill_depth_mm,
y = bill_length_mm)) +
geom_point() +
geom_smooth(method = "lm") +
annotate("text",
x = Inf, y = Inf, # This puts the text in the top-right corner of the plot.
label = paste0("Pearson R = ", # paste0() Combines (aka concatenates) pieces into one string (a string is one line of text or numeric values)
round(corr, 2)), # Add the correlation coefficient. Use round() to specify how many decimal places you want to show.
hjust = 1.1, vjust = 1.5)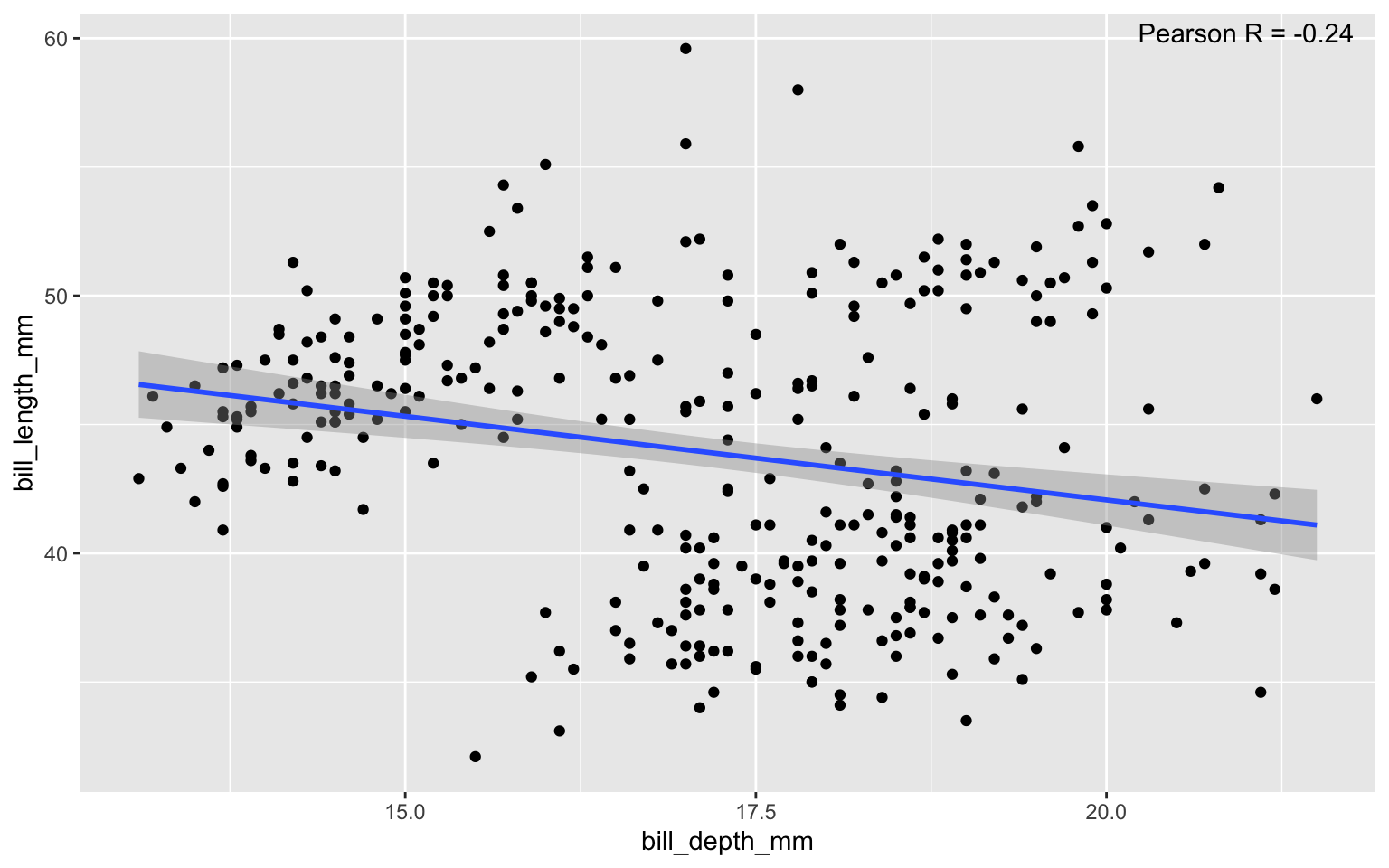
Side note: formatting plot elements
In the previous slide, we included annotations on the plot by adding the annotate() plot layer:
-
hjust: horizontal justification0= left0.5= center1= right
-
vjust: vertical justification0= bottom0.5= middle1= top
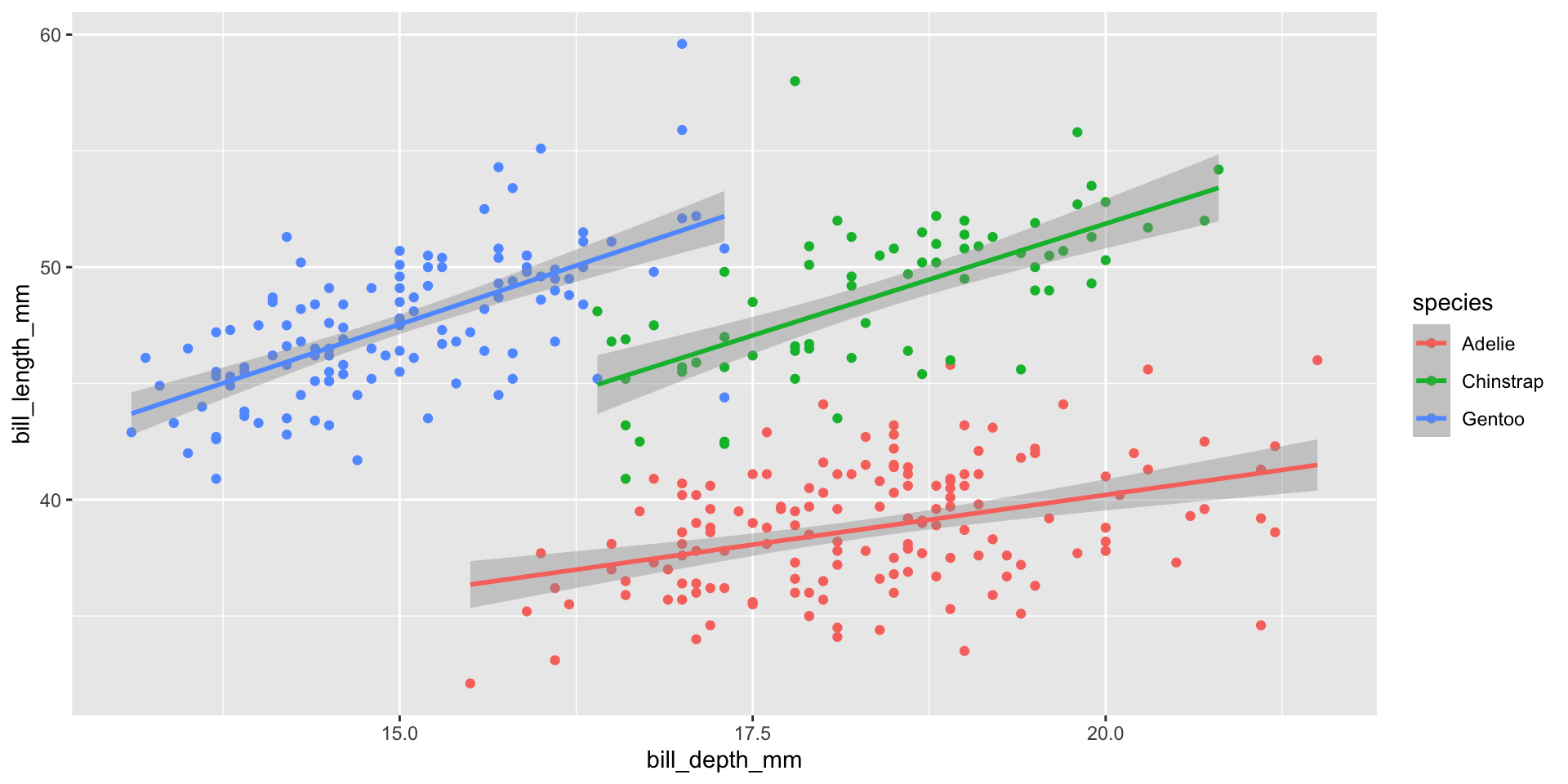
Relationships change based on groups within the data
Correlation coefficient for groups within data
- Relationships between variables might differ when we look at groups within the data.
- Use
group_by()beforesummarize()to calculate a separate Pearson’s R correlation coefficient for each group.
Scatter plots: add linear regression line and correlation coefficient
corr_by_species <- penguins |>
group_by(species) |>
summarize(
pearson_r = cor(bill_depth_mm, bill_length_mm, use = "complete.obs"),
x = max(bill_depth_mm, na.rm = TRUE),
y = max(bill_length_mm, na.rm = TRUE)
)
ggplot(penguins,
aes(x = bill_depth_mm,
y = bill_length_mm,
color = species)) +
geom_point() +
geom_smooth(method = "lm") +
geom_text(data = corr_by_species,
aes(x = x,
y = y,
label = paste0("r = ", round(pearson_r, 2))),
hjust = 1.1, vjust = 1.5)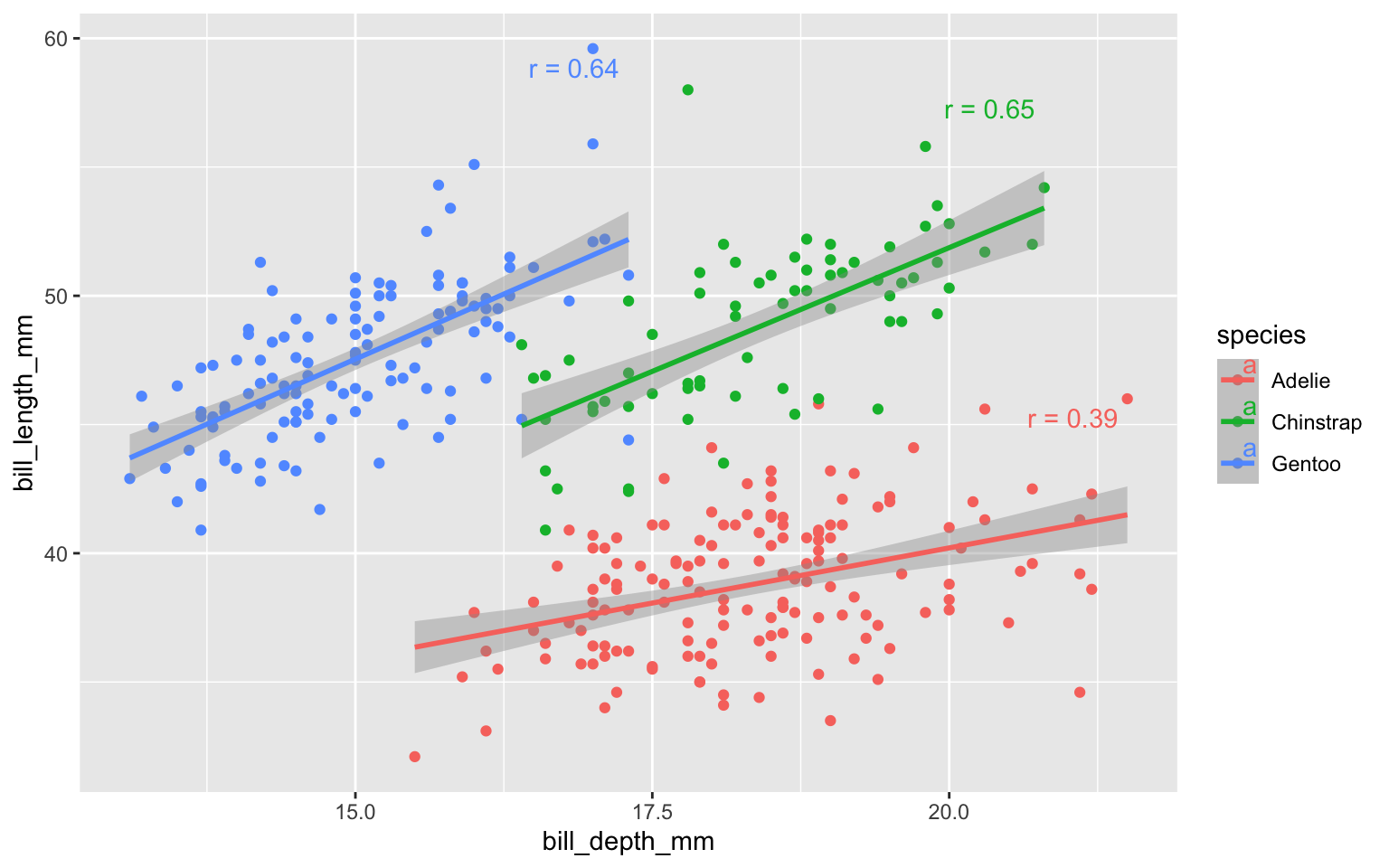
What did you notice about the linear regression lines?
Why did the linear relationship change?
Cross tabulations
Create cross tabulations using: count(), or group_by and summarize
Cross tabulations
- Compare how two variables relate by counting combinations of values.
Use count()
OR
group_by and summarize() with n()
- These produce a table showing how often each combination occurs
Examples
# Within each species, what share are on each island?
penguins |>
count(species, island) |>
group_by(species) |>
mutate(prop = n / sum(n))# A tibble: 5 × 4
# Groups: species [3]
species island n prop
<fct> <fct> <int> <dbl>
1 Adelie Biscoe 44 0.289
2 Adelie Dream 56 0.368
3 Adelie Torgersen 52 0.342
4 Chinstrap Dream 68 1
5 Gentoo Biscoe 124 1 # Within each island, what share is made up by each species?
penguins |>
count(species, island) |>
group_by(island) |>
mutate(prop = n / sum(n))# A tibble: 5 × 4
# Groups: island [3]
species island n prop
<fct> <fct> <int> <dbl>
1 Adelie Biscoe 44 0.262
2 Adelie Dream 56 0.452
3 Adelie Torgersen 52 1
4 Chinstrap Dream 68 0.548
5 Gentoo Biscoe 124 0.738